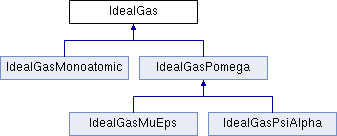
Abstract base class for an IdealGas evaluator.
More...
#include <IdealGas.h>
|
|
| IdealGas (int nIndep, const FluidMixture *, const FluidComponent *) |
| | Initialize and register to be used with excess functional fex in its fluidMixture.
|
| |
| virtual void | initState (const ScalarField *Vex, ScalarField *indep, double scale, double Elo=-DBL_MAX, double Ehi=+DBL_MAX) const =0 |
| |
|
virtual void | getDensities (const ScalarField *indep, ScalarField *N, vector3<> &P0) const =0 |
| | Given the independent variables indep, compute the site densities N and G=0 component of polarization density P.
|
| |
| virtual double | compute (const ScalarField *indep, const ScalarField *N, ScalarField *Phi_N, const double Nscale, double &Phi_Nscale) const =0 |
| |
| virtual void | convertGradients (const ScalarField *indep, const ScalarField *N, const ScalarField *Phi_N, const vector3<> &Phi_P0, ScalarField *Phi_indep, const double Nscale) const =0 |
| |
|
double | get_Nbulk () |
| |
|
void | overrideBulk (double Nbulk, double mu) |
| | Override the values for bulk density and chemical potential set by fluidMixture::initialize()
|
| |
|
|
double | Nbulk |
| | equilibirum density of this molecule in the bulk mixture
|
| |
|
double | mu |
| | chemical potential for this molecule
|
| |
|
double | corrPrefac |
| | prefactor for dipolar rotational correlations
|
| |
Abstract base class for an IdealGas evaluator.
◆ compute()
Return the ideal gas free energy PhiNI = T Int N - T S + (V-mu).N (where S is implementation dependent) and accumulate the gradients w.r.t the site densities Nscale is the factor by which the site densities/moments were scaled after getDensities() in order to implement fixed N / charge neutrality. Accumulate explicit gradients of PhiNI w.r.t Nscale in Phi_Nscale; the implicit dependence through N is handled by FluidMixture.
Implemented in IdealGasMonoatomic, and IdealGasPomega.
◆ convertGradients()
Compute Phi_indep, the total gradients w.r.t indep, given th egradients of the entire functional w.r.t site densities, Phi_N, and polarization density G=0 Phi_P0. Nscale will be the same as in compute()
Implemented in IdealGasMonoatomic, and IdealGasPomega.
◆ initState()
| virtual void IdealGas::initState |
( |
const ScalarField * |
Vex, |
|
|
ScalarField * |
indep, |
|
|
double |
scale, |
|
|
double |
Elo = -DBL_MAX, |
|
|
double |
Ehi = +DBL_MAX |
|
) |
| const |
|
pure virtual |
Create an initial guess for the indep, in presence of V and the extra potential Vex The initial guess is typically taken to be scale times what would generate the equilibrium ideal gas density upto caps Elo and Ehi on the molecule energy configurations considered. This would also be a good place to logPrintf useful statistics about the potential for debugging
Implemented in IdealGasPsiAlpha, IdealGasMonoatomic, and IdealGasPomega.
The documentation for this class was generated from the following file: